My main research areas are in applied probability and analysis. See below for more detailed descriptions of some previous and current projects.
Kinetic Theory and Grain Boundary Coarsening
Currently, I have been looking at how two dimensional network micro and macrostructures evolve under various types of coarsening. A well-studied method for coarsening is driven by mean-curvature flow. Several methods exist for understanding how networks and their statistics evolve, ranging from the direct simulation of networks (Potts models, level set methods, etc.) to kinetic theories which use mean field assumptions for expressing grain areas and topologies with systems of nonlinear advection reaction equations. Along with Govind Menon and Bob Pego, I’ve explored a happy medium which uses piecewise deterministic Markov processes (PDMPs), which allows us to approximate networks as stochastic dynamical systems, and also incorporate first order correlations between cells when constructing limiting kinetic equations. You can view our results here.
I’ve also taken a look at kinetic equations which model foams with edges that can break. Here’s a simulation of what the breaking process looks like over an initially ordered foam:
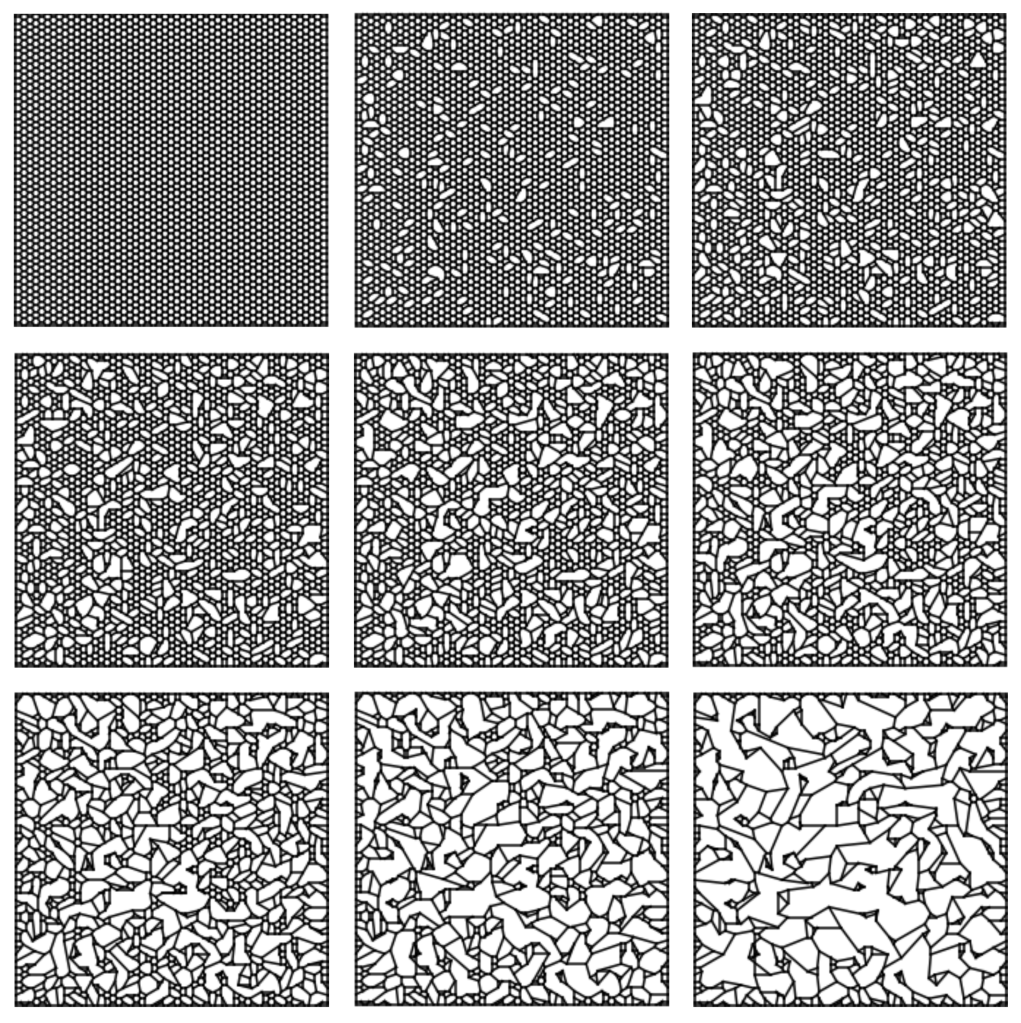

In this article, I develop a Markov process over the space of combinatorial embeddings and show that the associated kinetic equations are quite similar to the Smoluchowski coagulation equations, which are used for modeling coalescing particles.
To see how foams actually evolve in the wild, I’ve begun experiments with physical foams. Click here to see an animation showing what foam collapse looks like after applying a uniform heating of one of the plates.
Dynamics of Molecular Motors
In cells, a major issue is the transport of cellular cargo from one area to another. Here “cargo” can be genetic material, protein, or really any large molecule. We’re working at the microscale, so it’s technically possible for cargo to move through diffusion due to thermal fluctuations. For especially long cells (some neurons are over a meter long) , back of the envelope calculations give estimates on the order of years for traveling across the cell by diffusion alone.
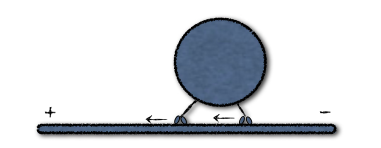
In the figure above, two molecular kinesin motors attached to a microtubule taxi a cargo from the minus to positive end. Scales are tiny, with a length of about 100 nm for the tether connecting the motor and cargo. At these sizes, thermal fluctuations kick in, along with some randomness involved with the mechano-chemical stepping process for actively moving the motors. Modeling the position of the two motors and cargo as a continuous process, the governing equations take the form of a system of nonlinear stochastic differential equations (SDEs).
With Peter Kramer (RPI) and John Fricks (ASU), I have developed a state dependent switching model which using principles of stochastic averaging theory of stochastic differential equations to derive effective statistics of motor ensembles. You can see our results here.
Self-Assembly in Nanotechnology
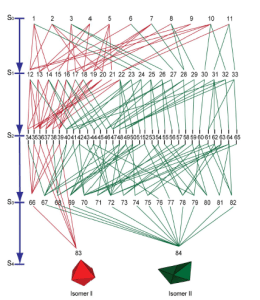
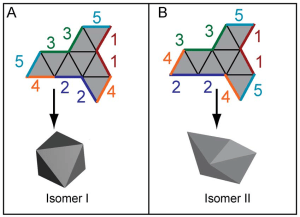
My first project at Brown was on the optimality of self-assembling nets. In collaboration with scientists from the Gracias Lab at Johns Hopkins University, we defined a gluing algorithm to create a configuration space of folding states. Viewed as a Markov chain, we derived a set of birth death equations which predict prevalent pathways that minimize defects.
Left: The path space for an octahedron and its non-convex cousin: the “boat”. Right: a single net with multiple end states.